Eukaryotic Chromosome Structure
DNA Structure, Replication and Eukaryotic Chromatin Structure Overheads
DNA Structure, Replication and Eukaryotic Chromatin Structure WWW Links
Eukaryotic Chromosome Structure
The length of DNA in the nucleus is far greater than the size of the compartment in which it is contained. To fit into this compartment the DNA has to be condensed in some manner. The degree to which DNA is condensed is expressed as its packing ratio.
Packing ratio - the length of DNA divided by the length into which it is packaged
For example, the shortest human chromosome contains 4.6 x 107 bp of DNA (about 10 times the genome size of E. coli). This is equivalent to 14,000 µm of extended DNA. In its most condensed state during mitosis, the chromosome is about 2 µm long. This gives a packing ratio of 7000 (14,000/2).
To achieve the overall packing ratio, DNA is not packaged directly into final structure of chromatin. Instead, it contains several hierarchies of organization. The first level of packing is achieved by the winding of DNA around a protein core to produce a "bead-like" structure called a nucleosome. This gives a packing ratio of about 6. This structure is invariant in both the euchromatin and heterochromatin of all chromosomes. The second level of packing is the coiling of beads in a helical structure called the 30 nm fiber that is found in both interphase chromatin and mitotic chromosomes. This structure increases the packing ratio to about 40. The final packaging occurs when the fiber is organized in loops, scaffolds and domains that give a final packing ratio of about 1000 in interphase chromosomes and about 10,000 in mitotic chromosomes.
Eukaryotic chromosomes consist of a DNA-protein complex that is organized in a compact manner which permits the large amount of DNA to be stored in the nucleus of the cell. The subunit designation of the chromosome is chromatin. The fundamental unit of chromatin is the nucleosome.
Chromatin - the unit of analysis of the chromosome; chromatin reflects the general structure of the chromosome but is not unique to any particular chromosome
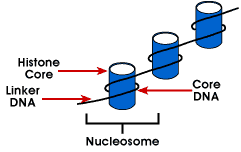
Nucleosome - simplest packaging structure of DNA that is found in all eukaryotic chromosomes; DNA is wrapped around an octamer of small basic proteins called histones; 146 bp is wrapped around the core and the remaining bases link to the next nucleosome; this structure causes negative supercoiling
The nucleosome consists of about 200 bp wrapped around a histone octamer that contains two copies of histone proteins H2A, H2B, H3 and H4. These are known as the core histones. Histones are basic proteins that have an affinity for DNA and are the most abundant proteins associated with DNA. The amino acid sequence of these four histones is conserved suggesting a similar function for all.
The length of DNA that is associated with the nucleosome unit varies between species. But regardless of the size, two DNA components are involved. Core DNA is the DNA that is actually associated with the histone octamer. This value is invariant and is 146 base pairs. The core DNA forms two loops around the octamer, and this permits two regions that are 80 bp apart to be brought into close proximity. Thus, two sequences that are far apart can interact with the same regulatory protein to control gene expression. The DNA that is between each histone octamer is called the linker DNA and can vary in length from 8 to 114 base pairs. This variation is species specific, but variation in linker DNA length has also been associated with the developmental stage of the organism or specific regions of the genome.
The next level of organization of the chromatin is the 30 nm fiber. This appears to be a solenoid structure with about 6 nucleosomes per turn. This gives a packing ratio of 40, which means that every 1 µm along the axis contains 40 µm of DNA. The stability of this structure requires the presence of the last member of the histone gene family, histone H1. Because experiments that strip H1 from chromatin maintain the nucleosome, but not the 30 nm structure, it was concluded that H1 is important for the stabilization of the 30 nm structure.
The final level of packaging is characterized by the 700 nm structure seen in the metaphase chromosome. The condensed piece of chromatin has a characteristic scaffolding structure that can be detected in metaphase chromosomes. This appears to be the result of extensive looping of the DNA in the chromosome.
The last definitions that need to be presented are euchromatin and heterochromatin. When chromosomes are stained with dyes, they appear to have alternating lightly and darkly stained regions. The lightly-stained regions are euchromatin and contain single-copy, genetically-active DNA. The darkly-stained regions are heterochromatin and contain repetitive sequences that are genetically inactive.
Centromeres and Telomeres
Centromeres and telomeres are two essential features of all eukaryotic chromosomes. Each provide a unique function that is absolutely necessary for the stability of the chromosome. Centromeres are required for the segregation of the centromere during me iosis and mitosis, and teleomeres provide terminal stability to the chromosome and ensure its survival.
Centromeres are those condensed regions within the chromosome that are responsible for the accurate segregation of the replicated chromosome during mitosis and meiosis. When chromosomes are stained they typically show a dark-stained region that is the centromere. During mitosis, the centromere that is shared by the sister chromatids must divide so that the chromatids can migrate to opposite poles of the cell. On the other hand, during the first meiotic division the centromere of sister chromatids must remain intact, whereas during meiosis II they must act as they do during mitosis. Therefore the centromere is an important component of chromosome structure and segregation.
Within the centromere region, most species have several locations where spindle fibers attach, and these sites consist of DNA as well as protein. The actual location where the attachment occurs is called the kinetochore and is composed of both DNA and protein. The DNA sequence within these regions is called CEN DNA. Because CEN DNA can be moved from one chromosome to another and still provide the chromosome with the ability to segregate, these sequences must not provide any other function.
Typically CEN DNA is about 120 base pairs long and consists of several sub-domains, CDE-I, CDE-II and CDE-III. Mutations in the first two sub-domains have no effect upon segregation, but a point mutation in the CDE-III sub-domain com pletely eliminates the ability of the centromere to function during chromosome segregation. Therefore CDE-III must be actively involved in the binding of the spindle fibers to the centromere.
The protein component of the kinetochore is only now being characterized. A complex of three proteins called Cbf-III binds to normal CDE-III regions but can not bind to a CDE-III region with a point mutation that prevents mitotic segregation. Fur thermore, mutants of the genes encoding the Cbf-III proteins also eliminates the ability for chromosomes to segregate during mitosis. Additional analyses of the DNA and protein components of the centromere are necessary to fully understand the mechanics of chromosome segregation.
Telomeres are the region of DNA at the end of the linear eukaryotic chromosome that are required for the replication and stability of the chromosome. McClintock recognized their special features when she noticed, that if two chromosomes were broken in a cell, the end of one could attach to the other and vice versa. What she never observed was the attachment of the broken end to the end of an unbroken chromosome. Thus the ends of broken chromosomes are sticky, whereas the normal end is not sticky, suggesting the ends of chromosomes have unique features. Usually, but not always, the telomeric DNA is heterochromatic and contains direct tandemly repeated sequences. The following table shows the repeat sequences of several species. These are often of the form (T/A)xGy where x is between 1 and 4 and y is greater than 1.
Telomere Repeat Sequences
| Species | Repeat Sequence |
|---|---|
| Arabidopsis | TTTAGGG |
| Human | TTAGGG |
| Oxytricha | TTTTGGGG |
| Slime Mold | TAGGG |
| Tetrahymena | TTGGGG |
| Trypanosome | TAGGG |
| Yeast | (TG)1-3TG2-3 |
Notice that the number of TG sequences and the number of cytosines in the yeast sequence varies. At least for yeast, it has been shown that different strains contain different lengths of teleomeres and that the length is under genetic control.
The primary difficulty with telomeres is the replication of the lagging strand. Because DNA synthesis requires a RNA template (that provides the free 3'-OH group) to prime DNA replication, and this template is eventually degraded, a short single-stranded region would be left at the end of the chromosome. This region would be susceptible to enzymes that degrade single-stranded DNA. The result would be that the length of the chromosome would be shortened after each division. But this is not seen.
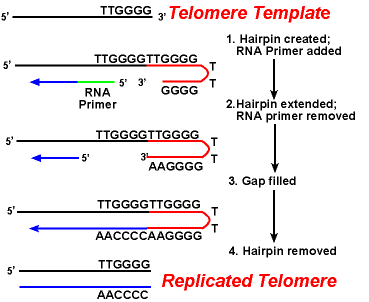
The action of the telomerase enzymes ensure that the ends of the lagging strands are replicated correctly. A well-studied system involves the Tetrahymena protozoa organism. The telomeres of this organism end in the sequence 5'-TTGGGG-3'. T he telomerase adds a series of 5'-TTGGGG-3' repeats to the ends of the lagging strand. A hairpin occurs when unusual base pairs between guanine residues in the repeat form. Next the RNA primer is removed, and the 5' end of the lagging strand can be used for DNA synthesis. Ligation occurs between the finished lagging strand and the hairpin. Finally, the hairpin is removed at the 5'-TTGGGG-3' repeat. Thus the end of the chromosome is faithfully replicated. The following figure shows these steps.
The Replication of Telomeres
Analysis of DNA Sequences in Eukaryotic Genomes
The technique that is used to determine the sequence complexity of any genome involves the denaturation and renaturation of DNA. DNA is denatured by heating which melts the H-bonds and renders the DNA single-stranded. If the DNA is rapidly cooled, the DNA remains single-stranded. But if the DNA is allowed to cool slowly, sequences that are complementary will find each other and eventually base pair again. The rate at which the DNA reanneals (another term for renature) is a function of the species from which the DNA was isolated. Below is a curve that is obtained from a simple genome.
The Y-axis is the percent of the DNA that remains single stranded. This is expressed as a ratio of the concentration of single-stranded DNA (C) to the total concentration of the starting DNA (Co). The X-axis is a log-scale of the product of the initial concentration of DNA (in moles/liter) multiplied by length of time the reaction proceeded (in seconds). The designation for this value is Cot and is called the "Cot" value. The curve itself is called a "Cot" curve. As can be seen the curve is rather smooth which indicates that reannealing occurs slowing but gradually over a period of time. One particular value that is useful is Cot½ , the Cot value where half of the DNA has reannealed.
Steps Involved in DNA Denaturation and Renaturation Experiments
1. Shear the DNA to a size of about 400 bp.
2. Denature the DNA by heating to 100oC.
3. Slowly cool and take samples at different time intervals.
4. Determine the % single-stranded DNA at each time point.
The shape of a "Cot" curve for a given species is a function of two factors:
- the size or complexity of the genome; and
- the amount of repetitive DNA within the genome
If we plot the "Cot" curves of the genome of three species such as bacteriophage lambda, E. coli and yeast we will see that they have the same shape, but the Cot½ of the yeast will be largest, E. coli next and lambda smallest. Physically, the larger the genome size the longer it will take for any one sequence to encounter its complementary sequence in the solution. This is because two complementary sequences must encounter each other before they can pair. The more complex the genome, that is the more unique sequences that are available, the longer it will take for any two complementary sequences to encounter each other and pair. Given similar concentrations in solution, it will then take a more complex species longer to reach Cot½ .
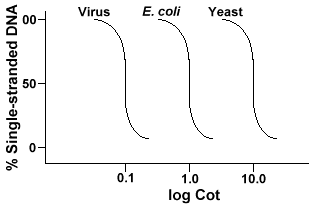
Repeated DNA sequences, DNA sequences that are found more than once in the genome of the species, have distinctive effects on "Cot" curves. If a specific sequence is represented twice in the genome it will have two complementary sequences to pair with and as such will have a Cot value half as large as a sequence represented only once in the genome.
Eukaryotic genomes actually have a wide array of sequences that are represented at different levels of repetition. Single copy sequences are found once or a few times in the genome. Many of the sequences which encode functional genes fall into this class. Middle repetitive DNA are found from 10s - 1000 times in the genome. Examples of these would include rRNA and tRNA genes and storage proteins in plants such as corn. Middle repetitive DNA can vary from 100-300 bp to 5000 bp and can be dispersed throughout the genome. The most abundant sequences are found in the highly repetitive DNA class. These sequences are found from 100,000 to 1 million times in the genome and can range in size from a few to several hundred bases in length. These sequences are found in regions of the chromosome such as heterochromatin, centromeres and telomeres and tend to be arranged as a tandem repeats. The following is an example of a tandemly repeated sequence:
ATTATA ATTATA ATTATA // ATTATA
Genomes that contain these different classes of sequences reanneal in a different manner than genomes with only single copy sequences. Instead of having a single smooth "Cot" curve, three distinct curves can be seen, each representing a different repetition class. The first sequences to reanneal are the highly repetitive sequences because so many copies of them exist in the genome, and because they have a low sequence complexity. The second portion of the genome to reanneal is the middle repetitive DNA, and the final portion to reanneal is the single copy DNA. The following diagram depicts the "Cot" curve for a "typical" eukaryotic genome
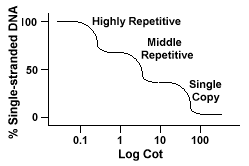
The following table gives the sequence distribution for selected species.
| Species | Sequence Distribution |
|---|---|
| Bacteria | 99.7% Single Copy |
| Mouse | 60% Single Copy 25% Middle Repetitive 10% Highly Repetitive |
| Human | 70% Single Copy 13% Middle Repetitive 8% Highly Repetitive |
| Cotton | 61% Single Copy 27% Middle Repetitive 8% Highly Repetitive |
| Corn | 30% Single Copy 40% Middle Repetitive 20% Highly Repetitive |
| Wheat | 10% Single Copy 83% Middle Repetitive 4% Highly Repetitive |
| Arabidopsis | 55% Single Copy 27% Middle Repetitive 10% Highly Repetitive |
Sequence Interspersion
Even though the genomes of higher organisms contain single copy, middle repetitive and highly repetitive DNA sequences, these sequences are not arranged similarly in all species. The prominent arrangement is called short period interspersion. This arrangement is characterized by repeated sequences 100-200 bp in length interspersed among single copy sequences that are 1000-2000 bp in length. This arrangement is found in animals, fungi and plants.
The second type of arrangement is long-period interspersion. This is characterized by 5000 bp stretches of repeated sequences interspersed within regions of 35,000 bp of single copy DNA. Drosophila is an example of a species with this uncommon sequence arrangement. In both cases, the repeated sequences are usually from the middle repetitive class. We discussed above where highly repetitive sequences are found.
Eukaryotic Chromosome Karyotype
Whereas bacteria only have a single chromosome, eukaryotic species have at least one pair of chromosomes. Most have more than one pair. Another relevant point is that eukaryotic chromosomes are detected only occur during cell division and not during all stages of the cell cycle. They are in their most condensed form during metaphase when the sister chromatids are attached. This is the primary stage when cytogenetic analysis is performed.
Each species is characterized by a karyotype. The karyotype is a description of the number of chromosomes in the normal diploid cell, as well as their size distribution. For example, the human chromosome has 23 pairs of chromosome, 22 somatic pairs and one pair of sex chromosomes. One important aspect of genetic research is correlating changes in the karyotype with changes in the phenotype of the individual.
One important aspect of genetics is correlating changes in karyotype with changes in phenotype. For example, humans that have an extra chromosome 21 have Down's syndrome. Insertions, deletions and changes in chromosome number can be detected by the skilled cytogeneticist, but correlating these with specific phenotypes is difficult.
The first discriminating parameter when developing a karyotype is the size and number of the chromosomes. Although this is useful, it does not provide enough detail to be begin the development of a correlation between structure and function (phenotype). To further distinguish among chromosomes, they are treated with a dye that stains the DNA in a reproducible manner. After staining, some of the regions are lightly stained and others are heavily stained. As described above, the lightly stained regions are called euchromatin, and the dark stained region is called heterochromatin. The current dye of chose is the Giemsa stain, and the resulting pattern is called the G-banding pattern.
C-Value Paradox
In addition to describing the genome of an organism by its number of chromosomes, it is also described by the amount of DNA in a haploid cell. This is usually expressed as the amount of DNA per haploid cell (usually expressed as picograms) or the number of kilobases per haploid cell and is called the C value. One immediate feature of eukaryotic organisms highlights a specific anomaly that was detected early in molecular research. Even though eukaryotic organisms appear to have 2-10 times as many genes as prokaryotes, they have many orders of magnitude more DNA in the cell. Furthermore, the amount of DNA per genome is correlated not with the presumed evolutionary complexity of a species. This is stated as the C value paradox: the amount of DNA in the haploid cell of an organism is not related to its evolutionary complexity. (Another important point to keep in mind is that there is no relationship between the number of chromosomes and the presumed evolutionary complexity of an organism.)
C Values of Organisms Used in Genetic Studies
| Species | Kilobases/haploid genome |
|---|---|
| E. coli |
4.5 x 103 |
| Human |
3.0 x 106 |
| Drosophila |
1.7 x 105 |
| Maize |
2.0 x 106 |
| Aribidopsis |
7.0 x 104 |
A dramatic example of the range of C values can be seen in the plant kingdom where Arabidopsis represents the low end and lily (1.0 x 10^8 kb/haploid genome) the high end of complexity. In weight terms this is 0.07 picograms per haploid Arabidopsis genome and 100 picograms per haploid lily genome.
Genome - the complete set of chromosomes inherited from a single parent; the complete DNA component of an individual; the definition often excludes organelles
Copyright © 1997. Phillip McClean