Estimating Distances Between Genes
Estimating Linkage From Three-Point Crosses
Recombination Involves Exchange Of Chromosomal Material
Measuring Linkage in Humans
Detecting and Measuring Linkage in Humans
As stated above, the detection of gene action in human is difficult because of the lack of controlled crosses and the low number of progeny. The same difficulties plague the determination of linkage in humans. But some linkages can be detected in humans using pedigree analysis. Let's look at the following pedigree and see if we can determine the linkage relationships.
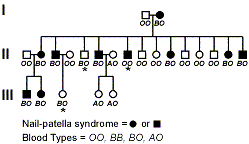
First, what can we determine about the inheritance of nail-patella syndrome? Clearly the disease is a dominant acting gene because affected individuals all had at least one parent with the disease. Secondly, can we determine the genotypes of the parents in generation I. The non-affected parent must be homozygous recessive for the disease. What about the affected parent in generation I. That parent must be heterozygous because offspring of this mating are homozygous recessive.
Next look at the blood type data. It is easier to genotype individuals for this phenotype because blood type is a codominant trait. Again the non-affected male is homozygous recessive whereas the affected female is heterozygous.
When we consider these two two genes, the parental mating in generation I is reminiscent of what type of cross? Well the male is homozygous recessive at both genes, and the female is heterozygous at both genes. This is equivalent to a testcross. If you remember, the test cross was used to determine linkage in all of the examples that were described above. So the next step is to determine if we are seeing independent assortment of the two genes or if the two genes appear to move as a block into the gametes.
By looking at the pedigree, we can see that in almost all cases, individuals with nail-patella syndrome also posses the B allele. This strongly suggests that the nail-patella and blood type genes are linked, and that the dominant allele responsible for the disease is in coupling with the B allele at the blood type locus.
As stated above, most but not all of the offspring show this linkage. Which of the offspring are the recombinants? If we look at generation II, the second male offspring of the marriage (II-5) does not have the disease but does contain the B allele. Also, in this generation, the fourth male offspring (II-8) has the disease, but does not have a B allele. In generation III, offspring (III-3) of the marriage of the first male in generation II has the B allele, but is not affected by the disease. So we can conclude that recombination has occurred between these two genes.
Now we would like to determine the linkage distance between the two genes. The original mating in genration I and the first two matings in generation II are test cross. The third mating in generation II is not informative because it involves the A allele which we are not following. We have a total of 16 offspring that are informative. Of these we determined that three were recombinant. As with all test crosses, this gives a genetic distance of 18.8 cM [100*(3/16)].
Copyright © 1998. Phillip McClean